 
We have adapted VMD to support EGL, permitting its use for parallel visualization without any dependency on windowing system software. Recently, it has become possible to eliminate the need for a running windowing system for visualization applications, through the use of the Embedded-system Graphics Library (EGL) in conjunction with the Vendor Neutral GL Dispatch Library (GLVND) that is part of current vendor-provided OpenGL drivers. VMD has previously relied on the OpenGL Extension to the X-Window System (GLX) to manage OpenGL rasterization on GPUs and with software-based rasterizers such as Mesa and OpenSWR. 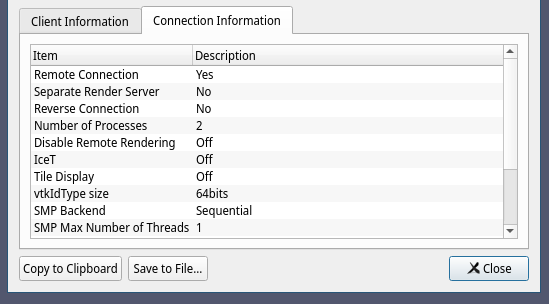 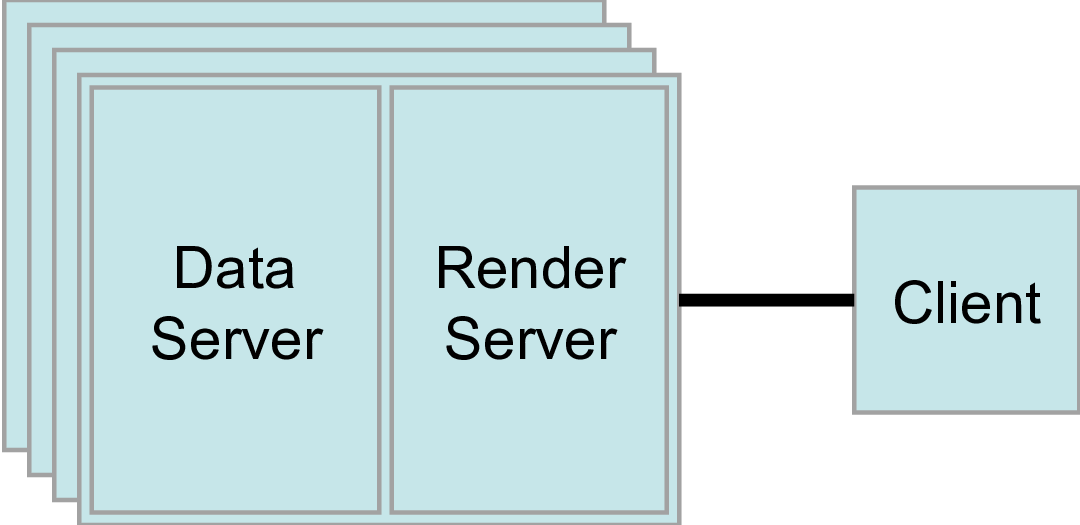
Unfortunately, glP remained a proprietary solution and it was orphaned when Sun exited the high performance graphics market. In 2004, Sun Microsystems proposed glP, an OpenGL extension for hardware accelerated rendering without the need for a windowing system. Until now it was not possible to develop vendor-neutral visualization applications that could take advantage of high performance OpenGL implementations except through windowing system-based software interfaces. VMD can be run as a conventional single-node interactive visualization application on remote visualization servers and clouds, but it also supports large scale parallel rendering on clusters and petascale supercomputers -,. This capability allows VMD to render and analyze MD trajectory data on-the-fly as it is produced. In the context of visualization on data center compute nodes, VMD supports in-situ visualization and computational steering of running MD simulations using a lightweight communication protocol originally developed for interactive haptic steering, , which is now supported by the popular NAMD, LAMMPS, GROMACS, and Terachem MD and quantum chemistry simulation tools. VMD is a widely used application for preparation, analysis, steering, and visualization of molecular dynamics simulations. Next-generation atomic-detail simulations of cellular complexes are expected to encompass billions of atoms, with attendant increases in visualization and analysis requirements. Routine visualization of the structure and dynamics of such biomolecular systems poses a formidable challenge, requiring parallel rendering with high performance graphics hardware and software. State-of-the-art molecular dynamics (MD) simulations of large viruses such as HIV, and photosynthetic membranes, contain tens to hundreds of millions of atoms, and produce terabytes of trajectory output, as do small-size but long-timescale (tens of microseconds) protein folding simulations. We expect that the techniques described here will be of broad benefit to many other visualization applications.Ĭontinuing advances in experimental imaging, structure refinement, and simulation of large biomolecular complexes such as viruses and photosynthetic organelles have created an increasing demand for high performance visualization. We then provide a brief evaluation of the use of EGL in VMD, with tests using developmental graphics drivers on conventional workstations and on Amazon EC2 G2 GPU-accelerated cloud instance types. We discuss the implementation of EGL support in VMD, a widely used molecular visualization application, and we outline benefits of the approach for molecular visualization tasks on petascale computers, clouds, and remote visualization servers. We outline the potential benefits of this approach in the context of visualization applications used in the cloud, on commodity clusters, and supercomputers. Recently, it has become possible for applications to use the Embedded-system Graphics Library (EGL) to eliminate the requirement for windowing system software on compute nodes, thereby eliminating a significant obstacle to broader use of high performance visualization applications. A significant challenge for deploying visualization software within clouds, clusters, and supercomputers involves the operating system software required to initialize and manage graphics acceleration hardware. It is therefore necessary to perform visualization tasks in-situ as the data are generated, or by running interactive remote visualization sessions and batch analyses co-located with direct access to high performance storage systems. Large scale molecular dynamics simulations produce terabytes of data that is impractical to transfer to remote facilities. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
